Regulatory genomics
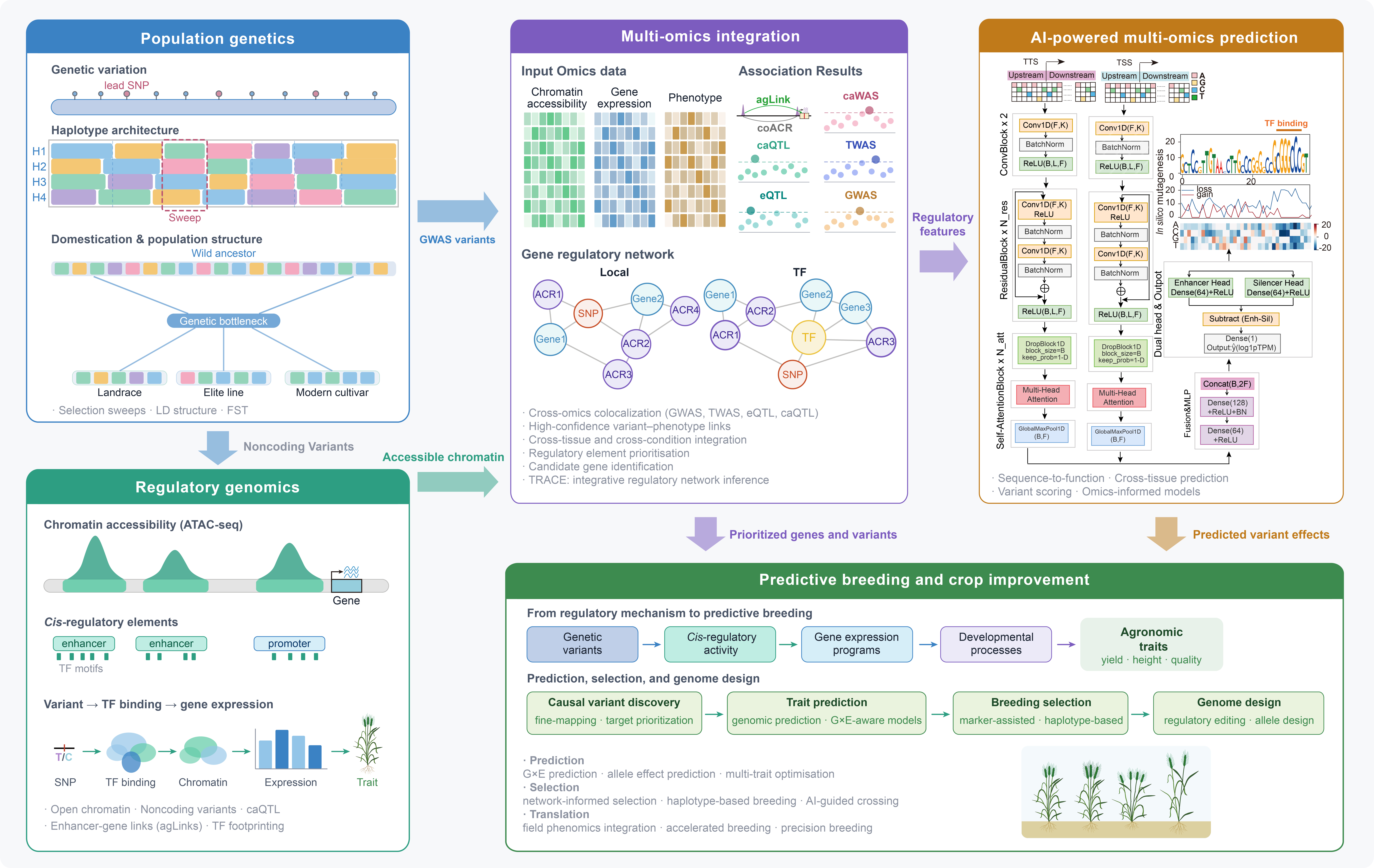
We investigate how noncoding regulatory variation shapes chromatin accessibility, transcriptional activity, and complex traits in wheat. A major goal of our work is to identify cis-regulatory elements, characterize their functional variation across populations, and understand how regulatory interactions contribute to developmental and agronomic phenotypes.