DeepWheat: predicting the effects of genomic variants on gene expression and regulatory activities across tissues and varieties in wheat using deep learning
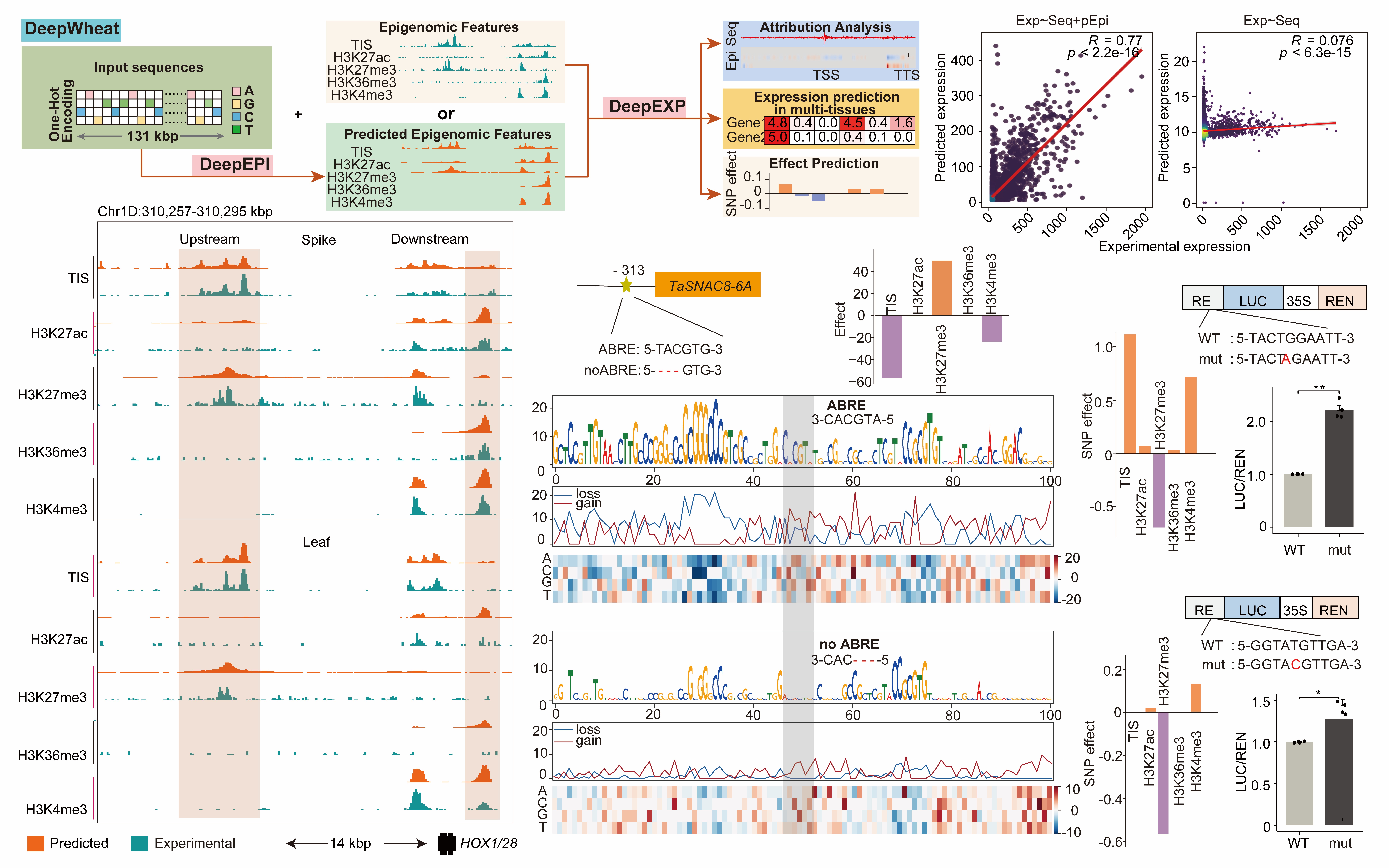
Spatiotemporal gene expression shapes key agronomic traits, yet tissue-specific prediction remains challenging in complex crops. We present DeepWheat, a broadly applicable deep learning framework comprising DeepEXP and DeepEPI, for accurate, tissue-specific gene expression prediction. DeepEXP integrates sequence and epigenomic features to predict gene expression (PCC 0.82–0.88), while DeepEPI predicts epigenomic maps from DNA sequence to support model transfer across varieties. Validations in five wheat cultivars confirm robustness and accuracy. DeepWheat also identifies regulatory variants with strong expression effects, enabling targeted cis-regulatory elements editing and offering a powerful tool for crop functional genomics and breeding.